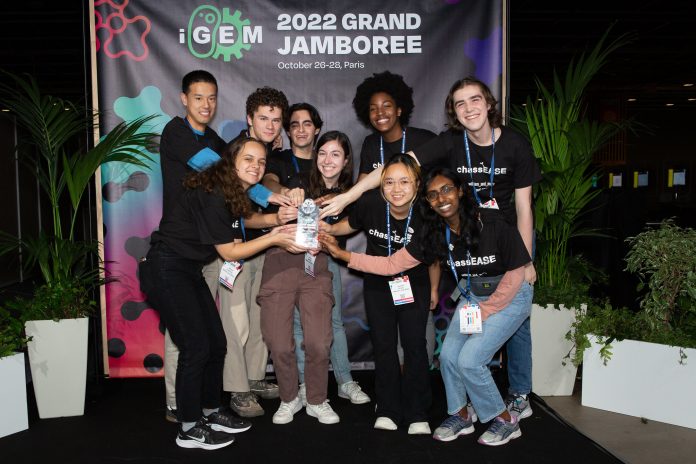
Oct. 26-28, the College of William and Mary’s International Genetically Engineered Machine team won best in the Software and AI development track at the 2022 Grand Jamboree held in Paris.
Competing against around 150 other teams, the College’s iGEM team also scored a gold medal, as well as nominations for Best Mathematical Model and Best Presentation.
iGEM is a global organization focused on fostering research in synthetic biology, or bioengineering, to create new biological systems. Research culminates annually at the Grand Jamboree for researchers at high school, undergraduate and graduate levels. This year, the College’s group was captained by biology majors Avery Bradley ’23 and Alana Thomas ’24. Faculty advisor and Chancellor Professor of Biology Margaret Saha and student advisor Beteel Abu-Ageel ’22 also supported the team in Paris.
This year’s team included Lin Fang ’24, Megan Fleeharty ’24, Walker Knapp ’25, Zhe Liu ’24, Krithika Layagala ’25, Diego Morandi ’24, Bjorn Shockey ’23 and Debby Zhong ’25.
The 2022 team chose to tackle the issue of fieldability, or a circuit’s ability to produce results in a natural environment. Their goal was to help researchers select an ideal chassis, or organism (usually bacteria) to house the circuit. They also hoped to further expand the applicability of synthetic biology beyond laboratory settings.
“Let’s say you’re a researcher and you built some system to bioremediate pollutants in the Chesapeake Bay,” Bradley said. “You can input specific kinds of parameters about the Chesapeake Bay into our program, and it will say ‘okay, you should use P. putida as your chassis,’ as a way to help researchers easily figure out the optimal chassis. Because ultimately, if the chassis can’t survive in that environment, your system isn’t going to work. It’s not going to produce any protein … it’s useless.”
The team began brainstorming for their competition concept in February 2022.
“We had our general theme of fieldability throughout the year, but we progressively tried to narrow down the scope,” Knapp said. “So we started with reviewing different areas in there … then, we shifted focus to modeling behavior of bacteria once you put them in the environment. And finally, we narrowed down … So it was this iterative process of narrowing down the scope of the project to focus on the core of a problem we could solve.”
Starting in late May and working throughout the summer, the team began research in both wetlab, or hands-on, experimental science, and drylab, or coding and math model-based, settings. In Bradley’s eyes, the dedicated research time lent the team a distinct focus.
“It’s fun because it’s a unique opportunity to really immerse yourself in the research that you don’t really get during the school year when you’re having to worry about classes and everything,” Bradley said. “I also really liked the team dynamic of iGEM. I think it’s different than other research labs because it was all of us constantly working together over the summer, collaborating and everything.”
“It’s fun because it’s a unique opportunity to really immerse yourself in the research that you don’t really get during the school year when you’re having to worry about classes and everything.”
The experimental phase continued throughout the beginning of the 2022 fall semester. The drylab team, composed of Knapp, Shockey and Liu, worked on data analysis and refining the software.
“We were mostly trying to analyze large amounts of metagenomic data for different bacteria cultures,” Knapp said. “What that data actually is is it’s people that have gone out and have taken samples from the environment … [they] get a full readout of the different types of bacteria and what ratios are present in that sample. So we take in all of that data on the order of 200,000 – 300,000 of those samples from different sources, databases, individual studies that we analyze, and we try to collect all of those together and sort of merge the different parameters. … and we try to all unify that so that we can be put into the rest of the software.”
The wetlab team worked concurrently on collecting samples to verify the software’s accuracy.
“We actually went out to the college woods, and we dug up some soil, and then we did a bunch of wetlab stuff on it to get the relative abundance of bacteria in the soil,” Fang said. “This way we can test our software, because if we put the same environmental parameters into our software, ideally, what’s spit out by the software should be the same as the sequencing result we obtained from the lab outside.”
Reaching out to professionals in the field of synthetic biology, the iGEM team supplemented the spaces in their research with expertise from other researchers.
“We ran into a little bit of difficulty with a few knowledge gaps that showed up on the math team,” Knapp said. “We have a wide diversity of majors and areas of expertise, but there were only three of us on the math team so we really, we couldn’t do everything and especially a lot of statistics topics and modeling topics. … But ultimately, we reached out to different experts in the field, different professors here, and we tried our best to reach … beyond the three of us to fill all those different gaps.”
In addition to completing their project, the College’s iGEM team went above and beyond, creating outreach materials that earned them a gold medal.
“So we do not just science, it’s so much: it’s collaboration, it’s multidisciplinary, it’s outreach, teaching the next generation about this incredibly cool field called syn bio.”
“We had one member of the team design a game called ReTerraforming Earth based on Terraforming Mars,” Saha said. “We took it to high schools, and they played the game and loved it and wanted copies of it … And we worked with the School of Ed, and we had high school students here as well, doing outreach. So we do not just science, it’s so much: it’s collaboration, it’s multidisciplinary, it’s outreach, teaching the next generation about this incredibly cool field called syn bio.”
Fang, Knapp and Bradley echoed Saha’s sentiments regarding the interdisciplinary nature of iGEM. For Knapp, the interdisciplinary approach was especially evident in the specifics of the project.
“We had to bring in a bunch of different people with advanced biology knowledge to inform the purpose of the project. … and then from that point, we had to work with different predictive models to generate all this data, so that’s where the computer scientists came in,” Knapp said. “Our math experts or physics people figure out the ways we can plug all this information into the computer and how to spit out useful information. From that point, we have people who are good at graphic design and communication to go and present this data in a cohesive manner that can be used by other people in the field. So it was really crucial for this project especially to have that wide range of different skill sets on the team.”
Most of the team flew out to Paris to present at the competition, which was structured, as Bradley put it, like a large-scale science fair, complete with booths for each group. Groups would give a larger talk, then a more private talk for the judges. For some, it was their first time presenting at such a large-scale event, but the supportive team dynamic and general environment reduced some of that stress.
“Teams are really willing to share their projects and we communicate a lot. And I feel like it was a really welcoming environment and a great learning experience. Because we learned so much from other teams: their design process, their wet lab, and their software techniques.”
“Fundamentally, it is a competition, but you don’t feel that tense competitive feel when you say, play a soccer game,” Fang said. “Teams are really willing to share their projects and we communicate a lot. And I feel like it was a really welcoming environment and a great learning experience. Because we learned so much from other teams: their design process, their wet lab, and their software techniques.”
For Fang, synthetic biology is inherently an ideal field for international collaboration.
“Even if you’re in, say, India, and you speak a completely different language, and that you’re in a different timezone, we all do synthetic biology in the standard way,” Fang said. “We can communicate and improve each other because of this uniform language of synthetic biology. This is why iGEM is so important and international, because of this modularity. That’s why we’re able to get insights and provide insights to different teams across the globe.”
Unified by this common language, Bradley and the team were inspired by the projects presented and look forward to the next Grand Jamboree and beyond.
“It was really powerful seeing what everybody did,” Bradley said. “iGEM is really big on solving global problems, and so it’s all these projects that are all solving global problems altogether being done by people my age. It was really fascinating, it really feels like the future of science.”